Abstract
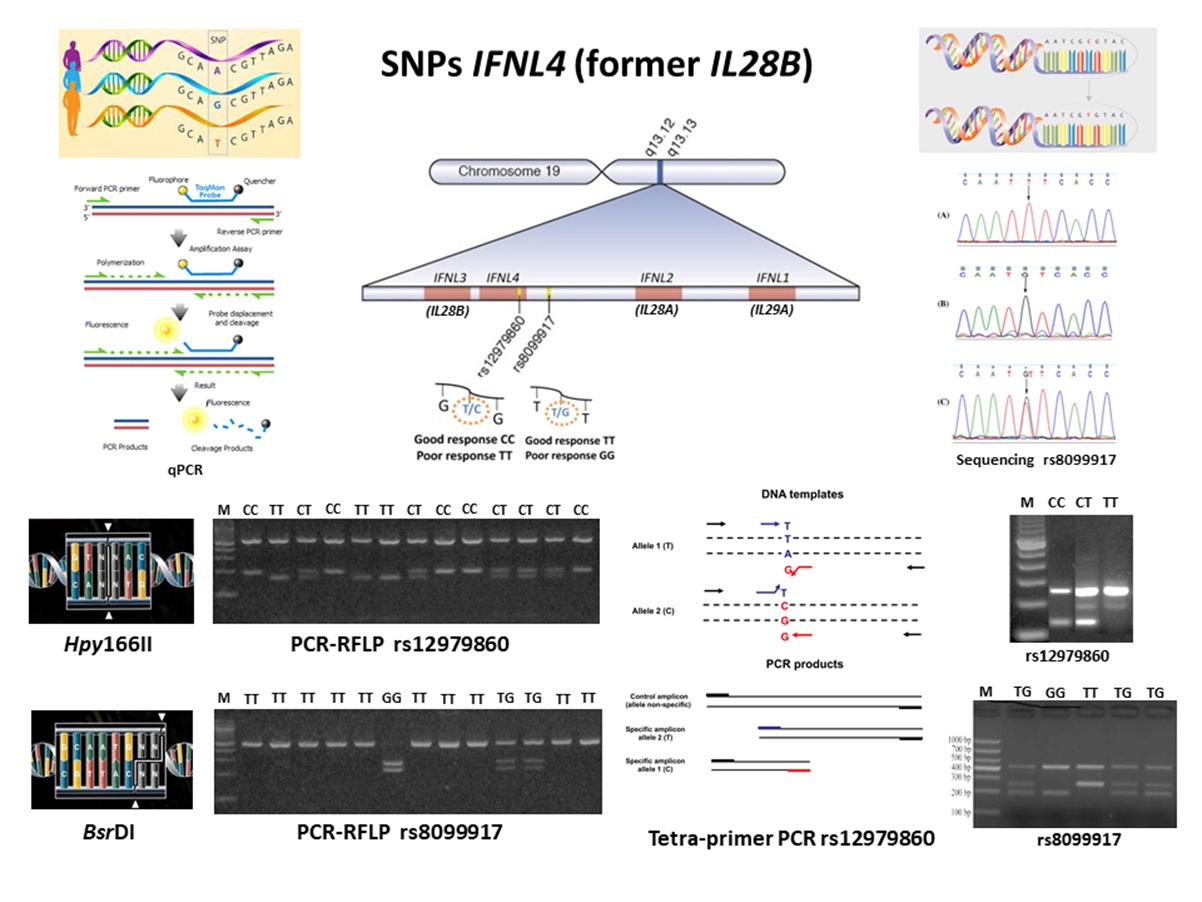
Single nucleotide polymorphisms (SNPs, rs12979860 e rs8099917) in the Interferon Lambda 4 gene (IFNL4, formerly IFNL3 and/or IL28B) has been associated with failure in the innate immune response, sustained virological response in hepatitis C, and HTLV-1-associated myelopathy (HAM) development. To search for these polymorphisms several methodologies can be employed, such as sequencing, real-time or quantitative polymerase chain reaction (qPCR), restriction fragment length polymorphism analysis in PCR products (PCR-RFLP), and tetra-primer PCR. The present study compared the performance of the tetra-primer PCR in relation to the PCR-RFLP, both optimized in the Research HTLV Laboratory of the Center of Immunology of Instituto Adolfo Lutz in São Paulo. One hundred DNA samples obtained from patients of STD/Aids Reference Centre in São Paulo, previously analyzed for IL28B SNPs by PCR-RFLP were selected for analysis, after confirming that they represent all IL28B SNPs patterns described in the literature. The results obtained showed concordance between the PCR-RFLP and the tetra-primer PCR SNPs results, and because of the low cost, easy to perform, and minor employment of biological specimen and reagents, the tetra- primer PCR is of choice to be used in routine.
References
1. Fang MZ, Jackson SS, O’Brien TR. IFNL4: Notable variants and associated phenotypes. Gene. 2020;730:144289. https://doi.org/10.1016/j.gene.2019.144289
2. Kotenko SV, Gallagher G, Baurin VV, Lewis-Antes A, Shen M, Shah NK et al. IFN – lambdas mediate antiviral protection through a distinct class II cytokine receptor complex. Nat Immunol. 2003;4(1):69-77. https://doi.org/10.1038/ni875
3. Sheppard P, Kindsvogel W, Xu W, Henderson K, Schlutsmeyer S, Whitmore TE et al. IL-28, IL-29 and their class II cytokine receptor IL-28R. Nat Immunol. 2003;4(1):63-8. https://doi.org/10.1038/ni873
4. Commins S, Steinke JW, Borish L. The extended IL-10 superfamily: IL-10, IL-19, IL-20, IL-22, IL-24, IL-26, IL-28, and IL-29. J Allergy Clin Immunol. 2008;121(5):1108-11. https://doi.org/10.1016/j.jaci.2008.02.026
5. Araujo ESA, Dahari H, Cotler JS, Layden TJ, Neumann AU, Melo CE et al. Pharmacodynamics of PEG-IFN alpha-2a and HCV response as a function of IL28B polymorphism in HIV/HCV co-infected patients. J Acquir Immune Defic Syndr. 2011;56(2):95-9. https://doi.org/10.1097/QAI.0b013e3182020596
6. Ramos JA, Ramos ALA, Hoffmann L, Perez RM, Coelho HSM, Ürményi TP. A single nucleotide polymorphism, rs129679860, in the IL28B locus is associated with the viral kinetics and a sustained virological response in a chronic, monoinfected hepatites C vírus genotype-1 Brazilian population treated with pegylated interferon-ribavirin. Mem Inst Oswaldo Cruz. 2012;107(7):888-92. https://doi.org/10.1590/S0074-02762012000700008
7. Assone T, Souza FV, Gaester KO, Fonseca LAM, Luiz OC, Malta F et al. IL28B gene polymorphism SNP rs8099917 genotype GG is associated with HTLV-1-associated myelopathy/tropical spastic paraparesis (HAM/TSP) in HTLV-1 carriers. PLoS Negl Trop Dis. 2014;8(9):e3199. https://doi.org/10.1371/journal.pntd.0003199
8. Assone T, Paiva A, Fonseca LAM, Casseb J. Genetic markers of the host in persons living with HTLV-1, HIV and HCV infections. Viruses. 2016;8(2):38. https://doi.org/10.3390/v8020038
9. Patel K, McHutchison J. Peginterferon alpha-2b: a new approach to improving response in hepatitis C patients. Expert Opin Pharmacother. 2001;2(8):1307-15. https://doi.org/10.1517/14656566.2.8.1
10. Ghany MG, Strader DB, Thomas DL, Seeff LB. Diagnosis, management, and treatment of hepatitis C: an update. Hepatology. 2009;49(4):1335-74. https://doi.org/10.1002/hep.22759
11. Ministério da Saúde. Protocolo clínico e diretrizes terapêuticas para hepatite viral C e coinfecções: manejo do paciente infectado cronicamente pelo genótipo 1 de HCV e fibrose avançada / Ministério da Saúde. Secretaria de Vigilância em Saúde. Departamento de DST, Aids e Hepatites Virais. – Brasília: Ministério da Saúde, 2013. Suplemento1.
12. Chen L, Borozan I, Feld J, Sun J, Tannis LL, Coltescu C et al. Hepatic gene expression discriminates responders and nonresponders in treatment of chronic hepatitis C viral infection. Gastroenterology. 2005;128(5):1437-44. https://doi.org/10.1053/j.gastro.2005.01.059
13. Kau A, Vermehren J, Sarrazin C. Treatment predictors of a sustained virologic response in hepatitis B and C. J Hepatol. 2008;49(4):634-51. https://doi.org/10.1016/j.jhep.2008.07.013
14. Ge D, Fellay J, Thompson AJ, Simon JS, Shianna KV, Urban TJ et al. Genetic variation in IL28B predicts hepatitis C treatment induced viral clearance. Nature. 2009; 461(7262):399-401. https://doi.org/10.1038/nature08309
15. Suppiah V, Moldovan M, Ahlenstiel G, Berg T, Weltman M, Abate ML et al. IL28B is associated with response to chronic hepatitis C interferon-alpha and ribavirin therapy. Nat Genet. 2009;41(10):1100-04. https://doi.org/10.1038/ng.447
16. Tanaka Y, Nishida N, Sugiyama M, Kurosaki M, Matsuura K, Sakamoto N et al. Genome-wide association of IL28B with response to pegylated interferon-α and ribavirin therapy for chronic hepatitis C. Nat Genet. 2009;41(10):1105-09. https://doi.org/10.1038/ng.449
17. Sharafi H, Pouryasin A, Alavian SM, Behnava B, Keshvari M, Mehrnoush L et al. Development and validation of a simple, rapid and inexpensive PCR-RFLP method for genotyping of common IL28B polymorphisms: a useful pharmacogenetic tool for prediction of hepatitis C treatment response. Hepat Mon. 2012;12(3):190-5. https://doi.org/10.5812/hepatmon.849
18. Thomas DL, Thio CL, Martin MP, Qi Y, Ge D, O’Huigin C et al. Genetic variation in IL28B and spontaneous clearance of hepatitis C virus. Nature. 2009;461(7265):798-802 https://doi.org/10.1038/nature08463
19. Rauch A, Kutalik Z, Descombes P, Cai T, Di Iulio J, Mueller T et al. Genetic variation in IL28B is associated with ch ronic hepatitis C and treatment failure: a genome-wide association study.Gastroenterology. 2010;138(4):1338-45, 1345,e1-7. https://doi.org/10.1053/j.gastro.2009.12.056
20. Galvão C. Vetores da doença de chagas no Brasil. Curitiba: Sociedade Brasileira de Zoologia, 2014. Available from: SciELO Books http://books.scielo.org/id/mw58j
21. Ye S, Dhillon S, Ke X, Collins AR, Day IN. An efficient procedure for genotyping single nucleotide polymorphisms. Nucleic Acids Res. 2001;29(17): E88-8. https://doi.org/10.1093/nar/29.17.e88
22. Galmozzi E, Del Menico B, Rametta R, Dongiovanni P, Fracanzani AL, Benedan L et al. A tetra-primer amplification refractory mutation system polymerase chain reaction for the evaluation of rs12979860 IL28B genotype. J Viral Hepat. 2011;18(9):628-30. https://doi.org/10.1111/j.1365-2893.2010.01349.x
23. Delvaux N, Costa VD, Costa MM, Lampe E. Comparison of four methods of genotyping IL28B polymorphisms in chronic hepatitis C patients. J Virol Methods. 2015;220:1-4. https://doi.org/10.1016/j.jviromet.2015.04.001
24. Li W, Zeng Y, Wang J, Zhou B, Zhang J, Zhang H et al. Predicting sustained viral response to hepatitis C using a rapid and simple IL28B rs8099917 genotyping assay. Antiviral Res. 2012;94(1):54-6. https://doi.org/10.1016/j.antiviral.2012.02.007
25. Moreira S, Garcia RFL, Gutberlet A, Bertol BC, Ferreira LE, Pinho MSL et al. A straightforward genotyping of the relevant IL28B SNPs for the prediction of hepatitis C treatment outcome. J Virol Methods. 2012;184(1-2):93-7. https://doi.org/10.1016/j.jviromet.2012.05.024
26. Ferreira PRA, Santos C, Côrtes R, Reis A, Tenore SB, Silva MH et al. Association between IL28B gene polymorphisms and sustained virological response in patients coinfected with HCV and HIV in Brazil. J Antimicrob Chemother. 2012;67(20):509-10. https://doi.org/10.1093/jac/dkr488
27. Melis R, Fauron C, McMillin G, Lyon E, Shirts B, Hubley LM et al. Simultaneous genotyping of rs12979860 and rs8099917 variants near the IL28B locus associated with HCV clearance and treatment response. J Mol Diagn. 2011;13(4): 446–51. https://doi.org/10.1016/j.jmoldx.2011.03.008
28. Payungporn S, Tangkyvanich P, Jantaradsamee P, Theamboonlers A, Poovorawan Y. Simultaneous quantitation and genotyping of hepatitis B virus by real-time PCR and melting curve analysis. J Virol Methods. 2004;120(2):131-40. https://doi.org/10.1016/j.jviromet.2004.04.012
29. Garcia RFL, Moreira S, Ramos ALA, Ferreira LE, Mattos AA, Tovo CV et al. Interleukin 28B-related polymorphisms: a pathway for understanding hepatitis C virus infection? World J Gastroenterol. 2013;19(42):7399-404. https://doi.org/10.3748/wjg.v19.i42.7399
30. Fateh A, Aghasadeghi M, Siadat SD, Vaziri F, Sadeghi F, Fateh R et al. Comparison of three different methods for detection of IL28 rs12979860 polymorphisms as a predictor of treatment outcome in patients with hepatitis C virus. Osong Public Health Res Perspect. 2016;7(2):83-9. https://doi.org/ 10.1016/j.phrp.2015.11.004

This work is licensed under a Creative Commons Attribution 4.0 International License.
Copyright (c) 2023 Instituto Adolfo Lutz Journal